> CELL CONFINEMENT, A STEP CLOSER TO IN VIVO CELL MIGRATION
Cell migration plays an extremely important role in the dynamics of biological systems, ranging from unicellular to complex multi-cellular ones. This application note shows the use of the 4Dcell confinement technology as a tool to study cell migration in 2D and 3D geometries. In this particular case, such technology enabled the validation of a universal model that states the relation between cell speed and cell persistence (Maiuri et al.,2015).
> Compatible with high-resolution microscopy
> Understand the environment that surrounds your cells
> Keep your cells focused during migration
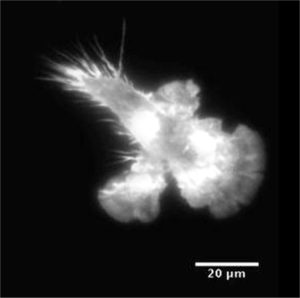

Mouse bone-marrow cell migrating in 2D while confined.
Maiuri et al.,2015
PRINCIPLE AND BENEFITS
The 4Dcell Confiner is a versatile device that enables the study of cell mechanics by applying well defined constrains to the cells. The method for confinement is based on the immobilization of cells between two parallel surfaces, enabling homogeneous and well-defined physical parameters, such as cell geometry and environment elasticity. Moreover, the confined cells can be imaged using high-resolution microscopy, as the device is optically transparent and maintains the cells at the focal plane.
EXPERIMENTAL SETUP
> Material needed
Confinement system: 4Dcell static confiner
Seeding substrate: glass bottom 6-well plate
Protein coating: fibronectin 25 µg/mL
> Cells used
Mouse bone-marrow cells (BMDCs)
> Protocol
Protocol for 2D migration with the static cell confiner
To reproduce the 2D migration assay, please proceed as follows:
Treat the 6-well plates with glass bottom (Matek) with a solution of 25 µg/mL of fibronectin for 60 min
Rinse the 6-well plates 3 times with PBS
Place your cells (50 000 cells per well) onto the surface and leave for 5 hours to adhere at 37oC
In the meantime, the static confiner, i.e., a 6-well lid containing the confinement pillars should be in medium in order to reach an equilibrium
After approximately 5 hours, the lid can be placed on top of the cells
After 2 hours, you can place the device in the microscope and acquire time-lapse imaging [1]
Protocol for 3D migration with the static cell confiner
To reproduce the 3D migration assay, please proceed as follows:
Prepare collagen gels using 1.6 mg/mL of bovine collagen (PureCol, Advanced BioMatrix)
Embed BMDCs in the polymerizing collagen
Immediately confine the gel between 2 glass surfaces spaced 5 µm apart with the cell confiner device [2]
After 20 min of collagen gelation at 37oC, add media to equilibrate the gel before beginning to image
Nota: as an alternative to the static cell confiner, the dynamic cell confiner can also be used to perform this type of experiments.
RESULTS
Studying cell migration can be a very challenging process, since common assays preclude long-term cell imaging with high-resolution, because ultimately cells can migrate in 3D.
In this particular case, the 4Dcell static confiner was used to study the migration of mouse bone-marrow cells in 2D and 3D geometries. In both cases BMDCs were confined between two parallel surfaces, 5 µm apart from each other, but while in the first case the cells were in a surface treated with fibronectin (A), in the second BMDCs were embedded in bovine collagen gel (B).
The following videos show the sequence of images retrieved to study the dynamics of cells migrating in the two different conditions.
Videos showing migration in conditions (A) and (B)
Cell dynamics retrieved from the analysis of each individual video:
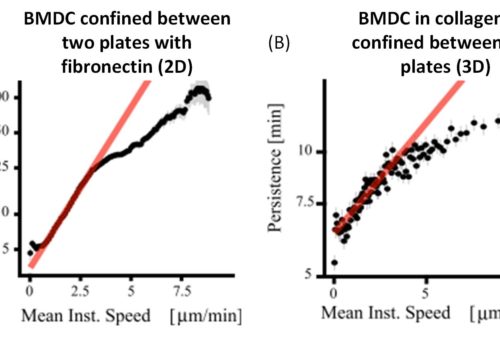
Figure 1: Plots resulting from the video analysis showing the relation between persistence and cell speed[3].
Based on the evidence found by Mauri et al., a correlation between the persistence time, t, and the mean instantaneous velocity, v, was found using distinct cell assays aiming to represent different conditions. This can be described by an exponential curve, t = Aeav, until the moment it saturates.
CONCLUSION
Closing the gap between in vivo and in vitro experiments leads to better-oriented research, and ultimately results in improvements in areas such as biomedicine.
Contrary to other assays where the physical environment of the cells is not fully controlled by the user, in this application note we showed the use of the 4DCell confiner to study the dynamics of BMDCs migrating in adhesive substrates and gels where the user controls the physical aspects of the experiments.
[1] In the particular case shown in this application note, a Zeiss AxioObserver microscope was used and the images were acquired using the phase contrast and fluorescence modes. In the case of fluorescence, the cell nuclei were labelled.
[2] The protocol is the same as for the 2D confinement. Confining the cells ensured that they remained in the same focal plane, allowing cell tracking during long periods of imaging.
[3] Persistence is the interval of time (Δt) that a cell moves in the same direction, and the mean instantaneous velocity is the length traveled by the cell divided by Δt. Red curves represent the exponential fit of the experimental data. The black dots represent the mean and the gray lines the standard error for the binned data on both axis.
REFERENCES
Actin flows induce a universal coupling between cell speed and cell persistence
P. Maiuri, R. Voituriez, et al. Cell. 2015,161(2), 374–386.
Fine control of nuclear confinement identifies a threshold deformation leading to lamina rupture and induction of specific genes
Le Berre M, Aubertin J, Piel M. Integr Biol (Camb).2012 Nov; 4 (11):1406-14.
