> STUDY IMMUNE CELL MIGRATION USING MICROCHANNELS
This protocol intends to provide information on how to analyze the migration of a large number of cells in conditions in which both the geometry and the biochemical nature of the environment are controlled, facilitating the normalization and reproducibility of independent experiments.
> Small sample volume and events multiplication
> Standardization of cell migration
> Compatible with fluoresence and high resolution imaging
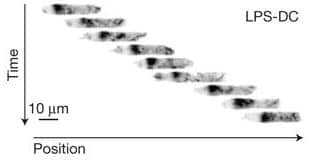

Displacement of one Dendritic cell along one channel. Vargas 2016.
PRINCIPLE AND BENEFITS
Inside channels and under confinement, cells have only two possibilities: move forward or backward. This simplified migration, restricted directionality, facilitates the automatic tracking of cells and the extraction of quantitative parameters to describe cell movement.
Microchannels are compatible with the use of fluorescent markers and are therefore suitable to study localization of intracellular organelles and structures during cell migration at high resolution. Finally, the surface of the channels can be functionalized with different substrates, allowing the control of the adhesive properties of the channels or the study of haptotaxis.
Migratory cells acquire the shape of the channels, adopting a tubular geometry that most likely reinforces their polarity.
APPLICATIONS
Chemotaxis and haptotaxis single cell analysis
3D cell culture systems
Drug discovery
Screening librairies generation
Immunoassays
EXPERIMENTAL SETUP
The protocol described here has been standardized using dendritic cells (DC), key actors in the immune system.
> Material needed
Small size microchannels: from 3 to 8 µm width
Protein coating: fibronectin 10 µg/mL
> Protocol
Preparation of 4Dcell microchannels dishes and cell plating
Please refer to the dedicated User Guide.
> Imaging
Delicately clean the glass external bottom surface.
After cell loading, video microscopy can be performed to follow cell migration. Figure 1 shows DC migrating in microchannels at an appropriate density to track the cells. Both phase contrast and Hoechst staining can be used for this purpose. The technique is also compatible with fluorescent confocal microscopy at higher resolution, and can be used to track organelles or cellular structures.
For time-lapse microscopy at 10X, choose a time frequency according to the expected cell speed, typically 2 min for dendritic cells (average migrating speed of 5 μm/min).
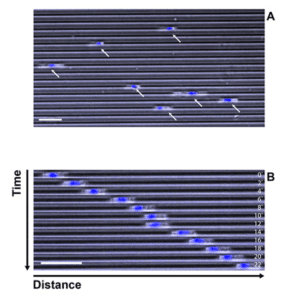

Figure 1: cells migrating along microchannels (5 µm width, 5 µm height). Phase contrast (grey) and nucleus staining (blue). A: single image extracted from movie. B: displacement of 1 DC along 1 channel (montage). Timescale 1 img /2 min. Bar: 50 µm
> Image analysis
Kymographs were generated from WT or Y27632 treated DCs.
NB: Y27632 is a well characterized inhibitor of cell contractility and migration, and was used to verify the ability of the system to detect changes in speed.
Several softwares can be used to track migrating cells. Some examples are described and compared in Hilsenbeck et al., 2016. Migration can be quantified thanks to phase contrast imaging (10x), or by fluorescence staining, which allows the tracking and analysis of migration with standard imaging softwares.
RESULTS
Dynamic of polymerized actin in migrating DCs expressing LifeActGFP (Figure 2)
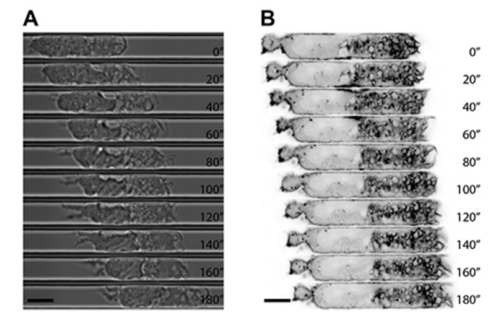
Figure 2: high resolution DCs migrating in microchannels (10 µm width, 5 µm height). A: Phase contrast images (1 img /2 min; montage). B: fluorescent imaging of cells expressing lifeAct (polymerized actin, in black) (1 img /2 min; montage) ; Bar: 10 µm
Quantification of cell motility in migrating DCs (Figure 3)
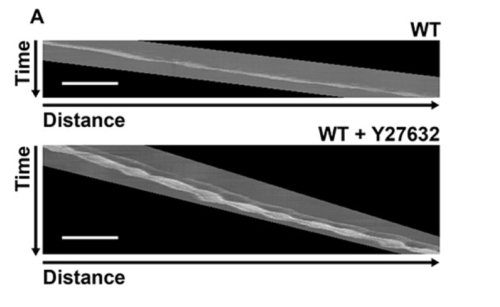
Figure 3: representative results. This illustrates the type of analysis that can be done from movies of cells migrating along channels. A: Kymographs (time lapse phase contrast, 10X, 1 img /2 min) of WT or Y27632 treated DCs. Bar : 100 µm; B: quantification of velocity, Mann-Whitney Stat. test.
CONCLUSION
Microchannels are a powerful tool to study the migratory properties of large numbers of cells in single experiments. This experimental system mimics the confined environmental constraints found in tissues by endogenous migratory cells. However, by forcing migration in a single direction, it facilitates automatic cell tracking and the extraction of measurables.
Compatible with fluorescence microscopy, microchannels are adapted to characterize speed of migration, cell behaviors, and organelles positioning during the different phases of cell motility. Extracellular matrix proteins, or channels’ geometry and size can also be modified, facilitating the study of the impact of physical constrains in cell migration.
This, together with a semi-automatized analysis, is compatible with medium throughput screenings for the discovery of new actors involved in the migration of cancer cells or for screening analysis of molecules involved in cell motility.
RELATED PUBLICATIONS
This application note is based on the great work of Pablo Vargas, in Matthieu Piel’s lab, UMR144 Institut Curie.
Study of cell migration in microfabricated channels
Vargas P, Piel M. et al., J Vis Exp. 2014
Software tools for single-cell tracking and quantification of cellular and molecular properties
Oliver Hilsenbeck et al., Nat. Biotech., 2016
